Research News
The Secret Life of Melons Revealed: "Jumping Sequences" May Alter Gene Expression
Researchers from the University of Tsukuba find that retrotransposons (a.k.a. "jumping sequences") may affect gene expression in melons
Tsukuba, Japan—On the surface, the humble melon may just look like a tasty treat to most. But researchers from Japan have found that this fruit has hidden depths: retrotransposons (sometimes called "jumping sequences") may change how genes are expressed.
In a study published recently in Communications Biology, researchers from the University of Tsukuba and the National Agriculture and Food Research Organization (NARO) have revealed that retrotransposons had a role in altering gene expression when melon genomes were diversifying, and may affect gene expression that induces fruit ripening.
Melons comprise one of the most economically important fruit crops globally. A special feature of melons is the coexistence of two fruit types: climacteric (which produce ethylene and exhibit a burst in cellular respiration as ripening begins), and non-climacteric. Ethylene is a plant hormone important to the regulation of climacteric fruit-ripening traits such as shelf life, which is of major economic importance.
"Because Harukei-3 melons produce ethylene during ripening, we wanted to look at ethylene-related gene expression in this type of melon," says lead author of the study Professor Hiroshi Ezura. "Harukei-3 produces an especially sweet fruit if grown in the right seasons. Because of its taste and attractive appearance, Harukei-3 has been used for a long time in Japan as a standard type for breeding high-grade muskmelon."
To examine ethylene-related gene expression, the researchers assembled the whole genome sequence of Harukei-3 by using third-generation nanopore sequencing paired with optical mapping and next-generation sequencing.
"We compared the genome of Harukei-3 with other melon genomes. Interestingly, we found that there are genome-wide presence/absence polymorphisms of retrotransposon-related sequences between melon accessions, and 160 (39%) were transcriptionally induced in post-harvest ripening fruit samples. They were also co-expressed with neighboring genes," explains Dr. Ryoichi Yano, senior author. "We also found that some retrotransposon-related sequences were transcribed when the plants were subjected to heat stress."
Retrotransposons are transposons (also referred to as "jumping sequences" because they can change their positions within a genome) with sequences similar to those of retroviruses.
"Our findings suggest that retrotransposons contributed to changes in gene expression patterns when melon genomes were diversifying. Retrotransposons may also affect gene expression that brings on fruit ripening," says Professor Ezura.
The Harukei-3 genome assembly, together with other data generated in this study, is available in the Melonet-DB database. Combined with future updates, this database will contribute to the functional genomic study of melons, especially reverse genetics using genome editing.
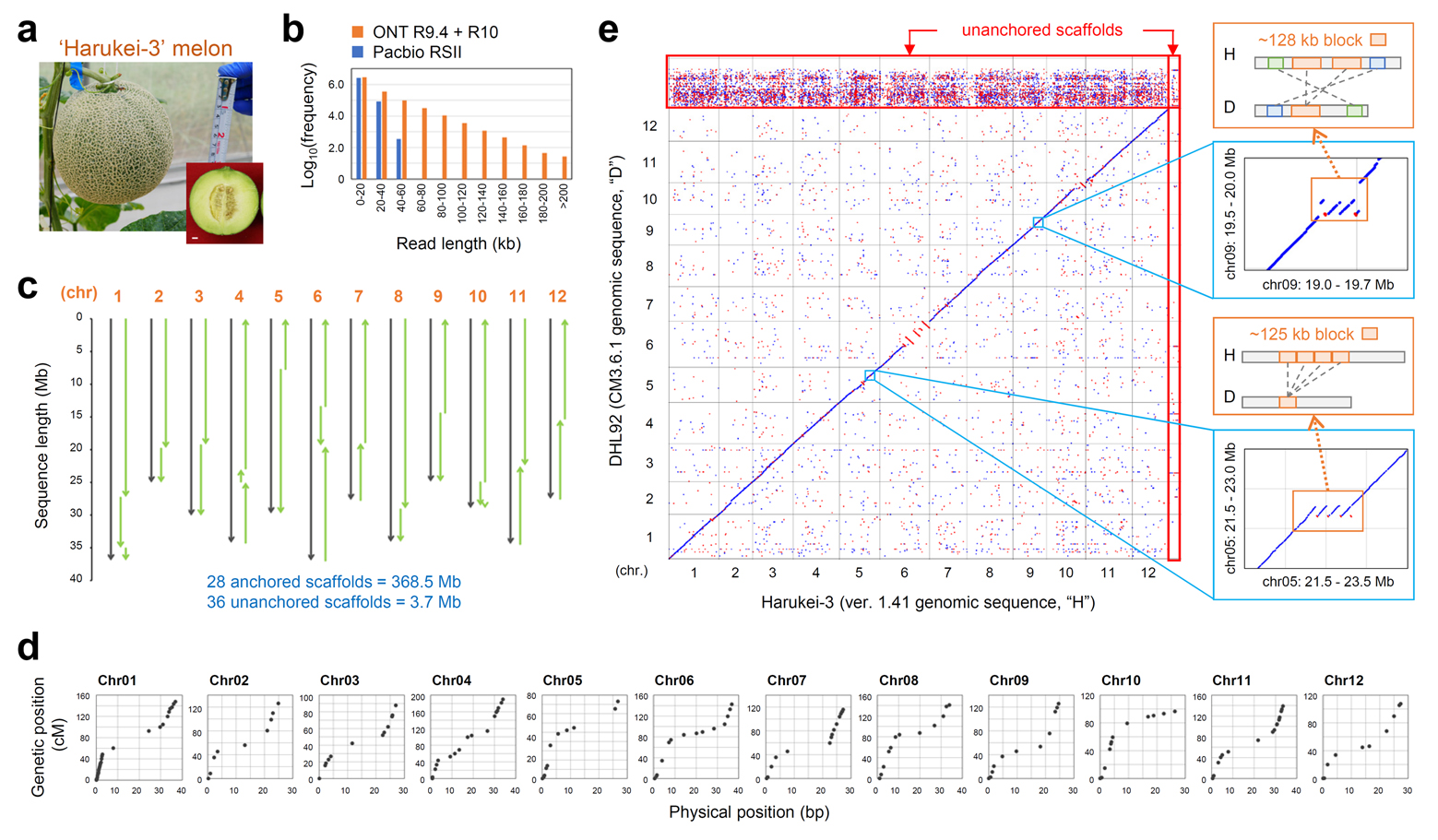
Figure: Whole genome assembly of Harukei-3 melon
Original Paper
The article, "Comparative genomics of muskmelon reveals a potential role for retrotransposons in the modification of gene expression," was published in Communications Biology at DOI: 10.1038/s42003-020-01172-0


