Research News
New Insights into Fermentation Enzyme Will Lower the Chemical Industry's Carbon Footprint
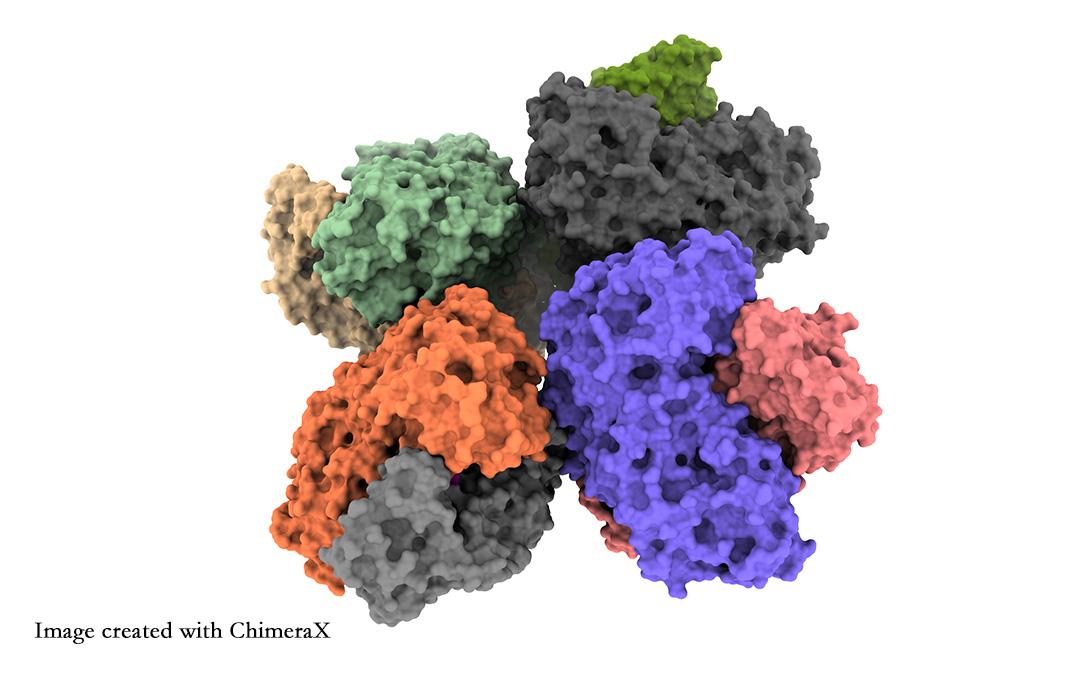
Researchers from the University of Tsukuba have obtained atomic-level insights into the structure of the phosphoketolase enzyme, which will help researchers optimize this enzyme for chemical feedstock synthesis
Tsukuba, Japan—Pharmaceuticals, plastics, and other industries use enzymes to help synthesize molecular feedstocks. Enzymes taken straight from microbes such as bacteria are often not optimal for industrial use; one issue is that they often do not survive the elevated temperatures that speed up a synthesis. Genetic engineering can help tailor enzymes for these purposes. Knowledge of the exact atom-by-atom structure of the original enzyme is important in understanding enzyme function in nature, thus providing insight as to how to optimize the genetic engineering of enzymes. However, X-ray crystallography, a common technique for determining an enzyme's structure as a critical step in this process, can unfortunately alter its structure as well.
A technique known as cryogenic electron microscopy (cryo-EM) can provide a similar level of structural detail to that of X-ray crystallography whilst retaining the native enzyme's structure. In fact, the 2017 Nobel Prize in Chemistry was awarded for using this technique to determine the structure of biological molecules. Now, in a study recently published in the Journal of Structural Biology, researchers from the University of Tsukuba and collaborating partners have used cryo-EM to determine the structure of the fermentation enzyme phosphoketolase. This work will facilitate genetic engineering of the enzyme for industrial syntheses.
"X-ray crystallography has revolutionized how researchers identify protein structures, but the development of alternative means that better reflect the structures seen in biology are invaluable," explains senior author Professor Kenji Iwasaki. "Our use of cryo-EM as an imaging tool has uncovered previously obscured structural detail in phosphoketolase that will directly benefit the chemical industry."
The researchers report two main findings. First, eight phosphoketolase units cluster together into one structure, known as an octamer. Second, they observed details of a chain of amino acids known as the QN-loop that may dictate whether the functional site of the enzyme is open or closed. This is a possible means of enhancing the chemical output of the enzyme.
X-ray crystallography obscures the structural detail provided by cryo-EM. The octamer was previously observed by X-ray crystallography but was thought to simply be a measurement artifact. Additionally, X-ray crystallography misses the open/closed structural details.
"Industry will now be able to correlate the function of phosphoketolase with its correct structure," says Iwasaki. "We expect that these insights will remind researchers that X-ray crystallography isn't necessarily the final word on enzyme structure; cryo-EM can offer valuable insights."
The results of this study are important for optimizing the performance of a fermentation enzyme that is useful for performing chemical syntheses in industry. By using enzyme structural insights to maximize the success of genetic engineering, feedstocks can be produced for pharmaceuticals, plastics, and other materials in an environmentally sustainable manner.
###
This research was partially supported by Platform Project for Supporting Drug Discovery and Life Science Research [Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)] from AMED under Grant Number JP 17am0101072 (to K.I. and N.M.).
Original Paper
The article, "High-resolution structure of phosphoketolase from Bifidobacterium longum determined by cryo-EM single-particle analysis," was published in the Journal of Structural Biology at DOI: 10.1016/j.jsb.2022.107842
Correspondence
Professor IWASAKI KenjiLife Science Center for Survival Dynamics, Tsukuba Advanced Research Alliance (TARA)
Related Link
Life Science Center for Survival Dynamics, Tsukuba Advanced Research Alliance (TARA)





